Frontier AI models have gotten genuinely good at science. Gemini 3.0 Pro, Claude Opus 4.5, and their peers can reason through complex multi-step problems, write real code, and engage with scientific concepts that felt like science fiction two years ago. But researchers deploying these models for serious work keep running into the same wall: raw intelligence isn't enough.
Ask a model about quantum circuit optimization and it can explain variational quantum eigensolvers in solid detail. Ask it to write Python code for molecular dynamics simulation and it'll produce something that compiles. But does it know your quantum computing group prefers Qiskit's native gates over transpiled circuits for benchmarking? Does it know your lab's conventions for LAMMPS input files, or the force field parameters you've validated over years of research?
This is the last mile problem of AI-powered scientific computing. It's exactly what Agent Skills were designed to solve.
The Intelligence Gap in Practice
The gap shows up everywhere.
A materials scientist asks for help with crystal structure prediction. The AI suggests using PyMatGen and the Materials Project API, technically correct. But it doesn't know that her group has a custom workflow for handling disordered alloys, or that they always cross-reference results against their internal DFT database before publication.
A quantum information researcher wants to simulate a variational circuit. The model writes valid PennyLane code, but it uses a hardware-agnostic approach when his lab specifically optimizes for IBM's superconducting qubit topology. The code runs; the results are suboptimal for his actual hardware.
A bioinformatician analyzing single-cell RNA-seq data gets a perfectly reasonable Scanpy pipeline. It doesn't use the QC thresholds her lab established through years of experience. It doesn't integrate with their downstream statistical methods. It doesn't know they export results to a specific electronic lab notebook format.
The model has intelligence; what it lacks is procedural knowledge and organizational context. This gap becomes even more pronounced in enterprise settings: pharmaceutical companies with proprietary ADMET protocols, national labs with classified simulation parameters, clinical research organizations with regulatory-compliant documentation standards. No matter how intelligent the underlying model becomes, it cannot absorb this institutional knowledge from training data alone.
Enter Agent Skills
In October 2025, Anthropic introduced Agent Skills - a deliberately simple approach to this problem.
At its core, a skill is just a folder with a SKILL.md file: instructions, examples, and optional supporting scripts. That's it. The simplicity is the point.
The key innovation is progressive disclosure. Rather than loading every possible piece of context into the model's context window (expensive, slow, and often counterproductive), skills allow agents to load information dynamically based on the task at hand.
At startup, the agent only sees the name and description of each available skill, a few sentences that help it understand when each skill might be relevant. When a task matches a skill's domain, the agent reads the full instructions. If those instructions reference additional files, the agent can read those too. This means a system can have hundreds of specialized capabilities without every conversation being buried in irrelevant context.
The architecture works like this:
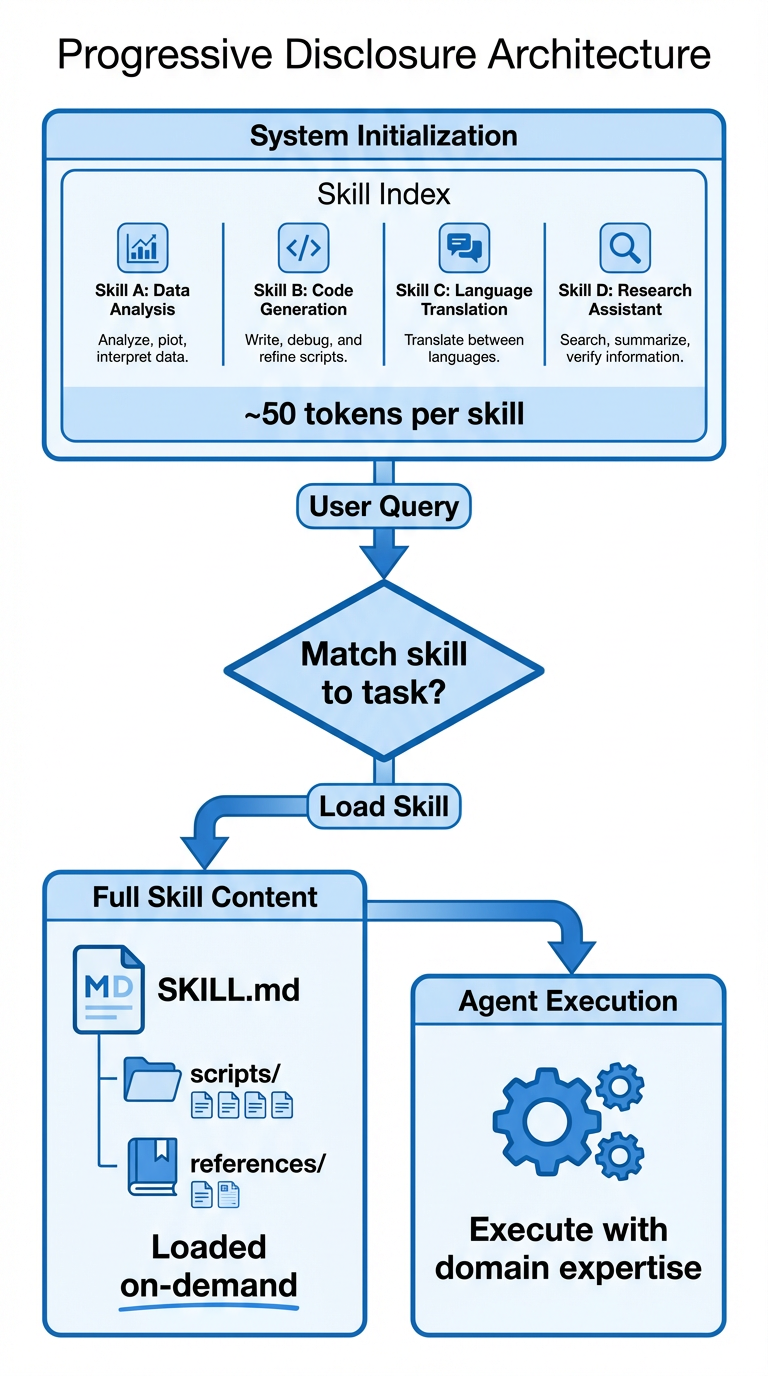
Figure 1: Progressive disclosure allows agents to maintain a compact skill index in memory while loading full skill content only when needed for a specific task.
In December 2025, Anthropic published Agent Skills as an open standard, so skills work across different AI platforms and tools. The specification is minimal by design: a folder with a SKILL.md file containing YAML frontmatter (just name and description) and Markdown instructions. This simplicity makes skills easy to write, version, share, and audit.
Scientific Agent Skills: Open Source Domain Expertise
When we at K-Dense saw what Agent Skills could do, we wanted to package and share what we'd already been building for K-Dense Web.
The result is Scientific Agent Skills, an open-source collection of 140 ready-to-use skills covering scientific computing across biology, chemistry, medicine, physics, and more.
The collection includes:
- 28+ Scientific Databases: Direct API access to OpenAlex, PubMed, bioRxiv, ChEMBL, UniProt, COSMIC, ClinicalTrials.gov, and more
- 55+ Python Packages: RDKit, Scanpy, PyTorch Lightning, scikit-learn, BioPython, BioServices, PennyLane, Qiskit, and others
- 15+ Scientific Integrations: Benchling, DNAnexus, LatchBio, OMERO, Protocols.io, and more
- 30+ Analysis & Communication Tools: Literature review, scientific writing, peer review, document processing, visualization
- 10+ Research & Clinical Tools: Hypothesis generation, grant writing, clinical decision support, regulatory compliance
Each skill includes comprehensive documentation, practical code examples, best practices, and integration guides. The entire collection is MIT-licensed, allowing commercial use and modification.
Skills vs MCPs: Complementary Approaches to Agent Capabilities
MCPs have become the de facto "API for AI" - a standard way for agents to interact with external services. At K-Dense, we use them. But skills and MCPs serve different purposes, and conflating them leads to confusion.
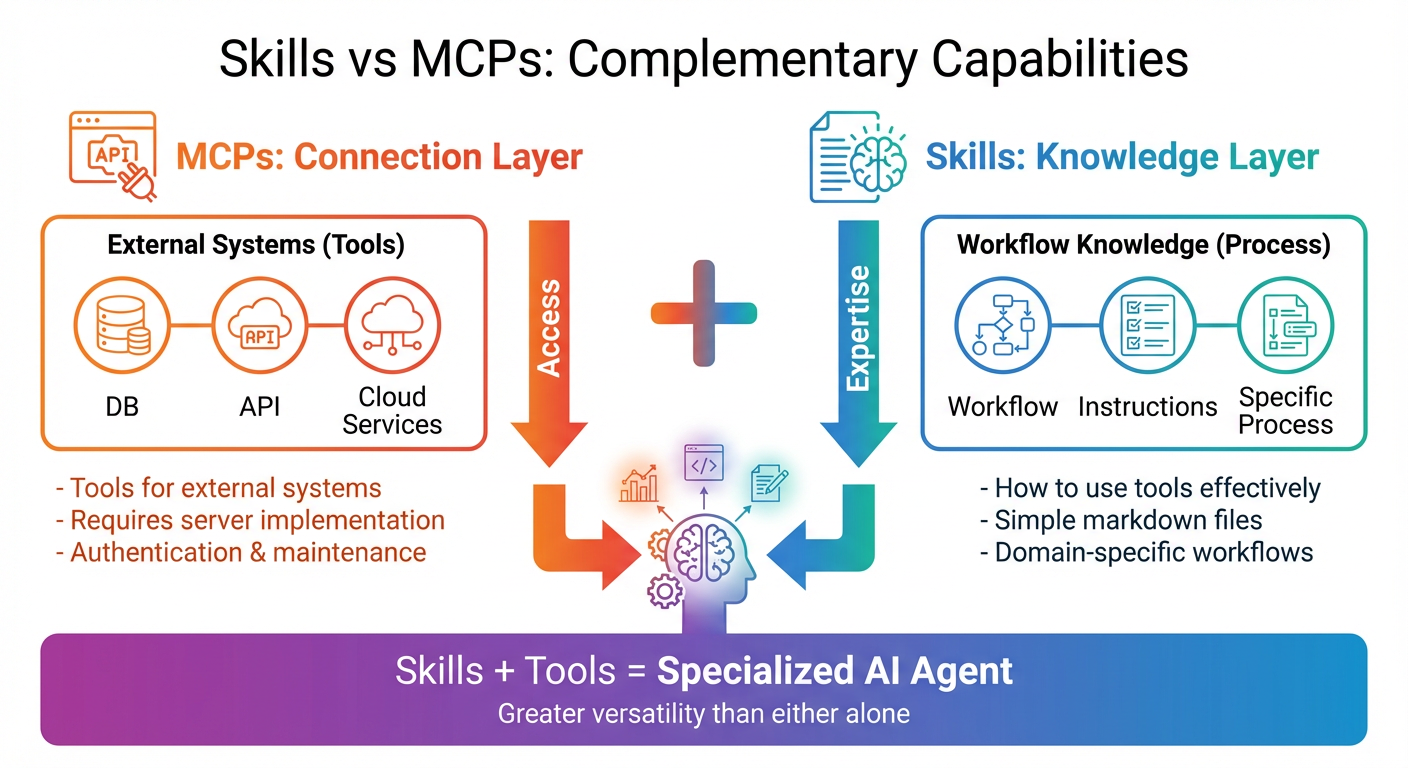
Figure 2: MCPs provide the connection layer to external systems, while skills provide the knowledge layer for using those systems effectively. Together, they create specialized AI agents with greater versatility than either approach alone.
MCPs are about connection. An MCP server provides an agent with access to an external service like a database, an API, or a computation engine. It's the bridge between the agent and the outside world. When you install an MCP for GitHub, the agent can read repositories, create issues, and manage pull requests. When you install an MCP for a database, the agent can query and modify data.
Skills are about knowledge. A skill teaches an agent how to use its capabilities effectively for a specific domain. It's not enough to have access to ChEMBL; you need to know how to formulate queries that return relevant molecular data for your drug discovery pipeline. It's not enough to be able to write Python code; you need to know the established workflows, best practices, and integration patterns for your specific analysis.
The relationship can be understood through a simple analogy. MCPs are the tools in a workshop: hammers, saws, drills, measuring instruments. Skills are the carpentry knowledge: understanding wood grain, joinery techniques, finishing methods, and design principles. A master carpenter needs both: tools without knowledge produce crude work, and knowledge without tools produces nothing at all.
This complementary nature means that skills plus fundamental tools provide far greater versatility than MCPs alone. Consider molecular docking for drug discovery. An MCP might provide access to AutoDock Vina for running docking calculations. But a skill provides the complete workflow: how to prepare ligands using RDKit, how to generate receptor structures from PDB files, how to configure grid boxes appropriately, how to interpret binding affinities, how to rank compounds for further investigation, and how to integrate results with your existing pipeline.
There's also a practical dimension. MCPs require real engineering investment - each server needs development, testing, maintenance, and updates as the underlying API changes. Authentication, error handling, the works. That burden adds up.
Skills are different. A skill is a Markdown file with optional supporting resources. A domain expert can write one in hours. Anyone who can read text can review, modify, or extend it. Version control is trivial. Sharing is copying a folder. For scientific work, where expertise sits with researchers rather than software engineers, that accessibility matters.
Context Efficiency and Cost
Token costs are real.
Progressive disclosure provides real efficiency gains - context loads only when needed. A system with 140 skills doesn't load 140 detailed instruction sets into every conversation. Instead, it loads a compact index of names and descriptions, typically consuming fewer than 10,000 tokens for the entire catalog. Only when a specific skill is activated does its full content enter the context window.
For cost-sensitive applications, this matters a lot. Consider a research platform handling thousands of queries per day. If every query loaded the full context for every possible capability, token consumption would be astronomical. With skills, a simple visualization question might load only a few hundred tokens, while a complex quantum chemistry workflow might load several thousand. The context scales with the task, not with the system's total capabilities.
There's also a quality dimension. Large context windows can actually degrade model performance. When an agent is overwhelmed with information, it struggles to identify the most relevant instructions. Skills help agents maintain precision and avoid "context confusion."
Open Source and the Scientific Community
We released Scientific Agent Skills under the MIT license so researchers, institutions, and companies can use, modify, and build on it without restriction.
This matches how scientific workflows actually evolve. A computational chemist writes a molecular docking skill for their drug discovery pipeline. A materials scientist at another institution adapts it for catalyst screening. A quantum computing researcher extends it for variational quantum chemistry. This is how methodological progress in science works - and it's what an open skill ecosystem enables.
The community response has been better than we expected. Contributors have submitted improvements, bug fixes, and entirely new skills covering domains we hadn't initially addressed. The collection has grown to 140+ skills, with new additions arriving regularly.
Maintaining a project of this scope requires sustained effort. We're committed to keeping skills updated as underlying tools evolve, integrating community contributions, and expanding coverage to new scientific domains. If you find these skills useful, we encourage you to contribute.
From Skills to Platform: K-Dense Web
Scientific Agent Skills is the foundation. K-Dense Web is what we built on top of it. The platform builds on everything in the open-source repository, and extends it with 200+ skills, cloud compute resources including GPUs and HPC, end-to-end research pipelines, and publication-ready outputs.
The key difference is integration. When you use skills directly with Claude Code or another agent environment, you're working with individual capabilities. K-Dense Web orchestrates these capabilities into coherent workflows, managing everything from data ingestion to final deliverables. Upload a dataset, describe your analysis objective, and the platform autonomously breaks down the task, selects appropriate methods, executes the analysis, and generates comprehensive reports with visualizations and statistical summaries.
That's the shift from AI as assistant to AI as executor. Traditional AI tools help you do work faster. K-Dense Web does the work, with your guidance. Tasks that would take a researcher days or weeks, such as comprehensive literature reviews, multi-method statistical analyses, machine learning pipeline development, complete in minutes.
Getting Started
There are a few ways to get started.
For Claude Code users: Install our skills directly as a plugin:
/plugin marketplace add K-Dense-AI/scientific-agent-skills
Then select and install the scientific-skills plugin. Once installed, simply mention a skill's domain in your conversation. Ask about quantum circuit simulation, materials property prediction, molecular docking, or literature review, and Claude will automatically activate the relevant skill.
For other environments: Clone the repository and integrate skills according to the agentskills.io specification. The format is simple enough that any agent framework with filesystem access can implement skill loading.
For the full experience: K-Dense Web provides everything in this repository plus additional capabilities, cloud infrastructure, and seamless workflow orchestration.
Separating procedural knowledge from model intelligence sounds like a small architectural choice. In practice, it's the difference between an AI that knows how science works in general and one that knows how your lab works specifically. That second thing is what researchers have been waiting for.
We've been surprised by how far a Markdown file goes. Write down your workflows, your tools, your conventions - and suddenly the model can help in a way that doesn't require you to rework everything afterward. That's the core insight. What the community builds from it is the interesting part.
Ready to transform your research with AI? Get started on K-Dense Web →
Questions? Email contact@k-dense.ai.
Related Resources:
