Walk into any working laboratory and what you see is not a wall of text.
You see a confocal stack of a fluorescent zebrafish heart on one monitor, a Western blot drying on the bench, an LC-MS chromatogram on a screen across the room, a lab notebook scrawled in pencil, and a voice memo from a PI about which condition to drop. You see slide decks with embedded gels, supplementary tables that nobody can find without scrolling, microscope GUIs with no API, whiteboard sketches of pathways, and seminar recordings that contain the most important sentence anyone said this month.
Science has always been multimodal. The question was when AI would finally be ready to meet it where it lives.
Today, NVIDIA introduced NVIDIA Nemotron 3 Nano Omni, an open multi-modal model with highest efficiency that powers sub-agents to complete tasks faster across vision, audio, and language. K-Dense is one of the partners evaluating Nemotron 3 Nano Omni, and we could not be more excited about what it means for agentic science.
This post is about why we believe a model like this matters, what it unlocks for the kind of work K-Dense Web does every day, and where this collaboration is heading next.
The Multimodal Reality of Doing Science
Most "AI for science" demos still pretend research is text. Read this paper. Summarize this protocol. Draft this section. That is a real and useful slice of the workflow, but it is a small one.
Here is a closer look at an actual day for a translational genomics group that uses K-Dense Web:
- Morning. 2,400 brightfield + DAPI tile images from an overnight high-content screen, organized by 384-well plate, plus a CellProfiler pipeline that almost works.
- Mid-morning. A 47-page preprint with seven figures, three of which contain panels the body text never describes in words.
- Lunch. A Zoom recording of a sponsor call, where a pharma partner asks the team to "reproduce that gel from Zhang et al.'s figure 4B but with our compound."
- Afternoon. A confocal instrument log in TIFF + JSON, an .mzML file from the mass spec, and a hand-drawn pathway sketch on a whiteboard photo.
- Evening. A Slack thread with three voice notes describing what the PI saw at the bench, ending with "we should also look at what happens if we drop the Mn²⁺."

A normal day in a working lab. None of these artifacts are reducible to text without losing what makes them useful.
A text-only AI co-scientist has a serious credibility problem in this room. It can summarize the paper, but cannot read the figures. It can take notes from the call, but cannot listen to the audio that contains the actual decision. It can write a Python script that loads images but cannot itself look at the images and tell you whether the segmentation worked.
The way scientists actually do work, and the way they actually communicate it to each other, is irreducibly multimodal. The way our agents reason has to follow.
Why a Pipeline of Specialists is Not Enough
The dominant pattern in 2025 was to bolt modalities together at the seam. A vision model labels the figure. A speech model transcribes the call. A document parser flattens the PDF into Markdown. A language model reads all those summaries side by side and tries to reason over the merged result.
This works until it does not. Each handoff loses fidelity:
- The vision model's caption omits the axis labels the language model needed.
- The transcript drops the inflection that turned a hedge into a deletion criterion.
- The document parser hands the LLM a flattened table that has lost its column headers.
- The audio summarizer collapses two minutes of "wait, that's actually weird" into "the team discussed the result."
Over a long-running agentic task, those small losses compound into wrong conclusions. We see this pattern often enough in benchmarking that we have stopped being surprised by it.
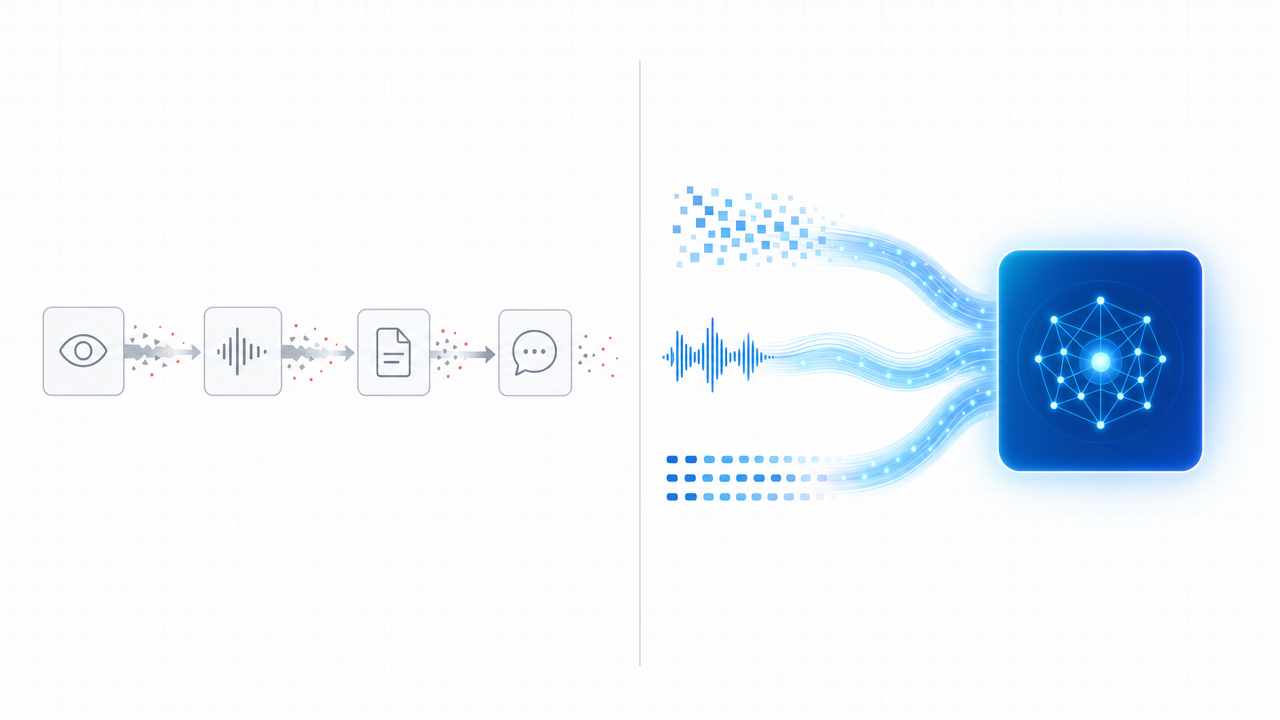
Left: a pipeline of specialists, leaking fidelity at every handoff. Right: a unified omni-modal model that carries pixels, audio, and tokens through a single reasoning stream.
Nemotron 3 Nano Omni was built explicitly to collapse this stack. By integrating vision and audio encoders directly with the language backbone, the model carries pixels, audio frames, and tokens through a single reasoning stream. There is no transcription step that throws away tone. No captioning step that throws away pixels. The model sees what the scientist sees, hears what the scientist heard, and reasons over all of it together.
The architectural choices that make this practical are also the right ones for science. The broader Nemotron 3 family is built on a hybrid Mamba-Transformer mixture-of-experts with long context, sparse activation, and granular reasoning-budget control at inference time, and independent third-party evaluators have repeatedly highlighted its efficiency at the frontier of open intelligence. For our workloads, which routinely involve thousands of pages of regulatory filings, large microscopy datasets, or multi-hour seminar recordings, that combination of long context and efficient sparse activation is exactly the right shape.
AI Co-Scientists Need to See the World as Scientists Do
We have written before about the gap between raw model intelligence and the procedural knowledge of a working lab. Agent Skills closed a big part of that gap on the language side, by letting agents load lab-specific workflows on demand. But there has been a second gap that text-and-skills alone could never close: perception.
A real co-scientist, the kind of person you actually want in the room when you are trying to interpret a result, is doing several things at once.
They are reading the experiment. The morphology in the well, the smear on the gel, the shape of the curve, the orientation of the bands. They are reading the room. The hesitation in someone's voice when they describe a result, the slide where the senior scientist visibly relaxed. They are reading the literature, but as scientists actually read it, by jumping straight to the figures and tables and circling back to the prose only when they need to.
A model that has only ever seen text has, on a basic perceptual level, never been in the room. It does not matter how clever the reasoning chain is. If the agent cannot see the panels, hear the call, or operate the instrument GUI, then on the most important parts of any scientific workflow, it is asking a colleague to describe the world for it.
Nemotron 3 Nano Omni is one of the first open models that meaningfully closes this perceptual gap while remaining efficient and deployable. That is why we are evaluating it. That is why we are collaborating with NVIDIA on it.
Three Agentic Patterns Where Omni-Modal Reasoning Changes the Game
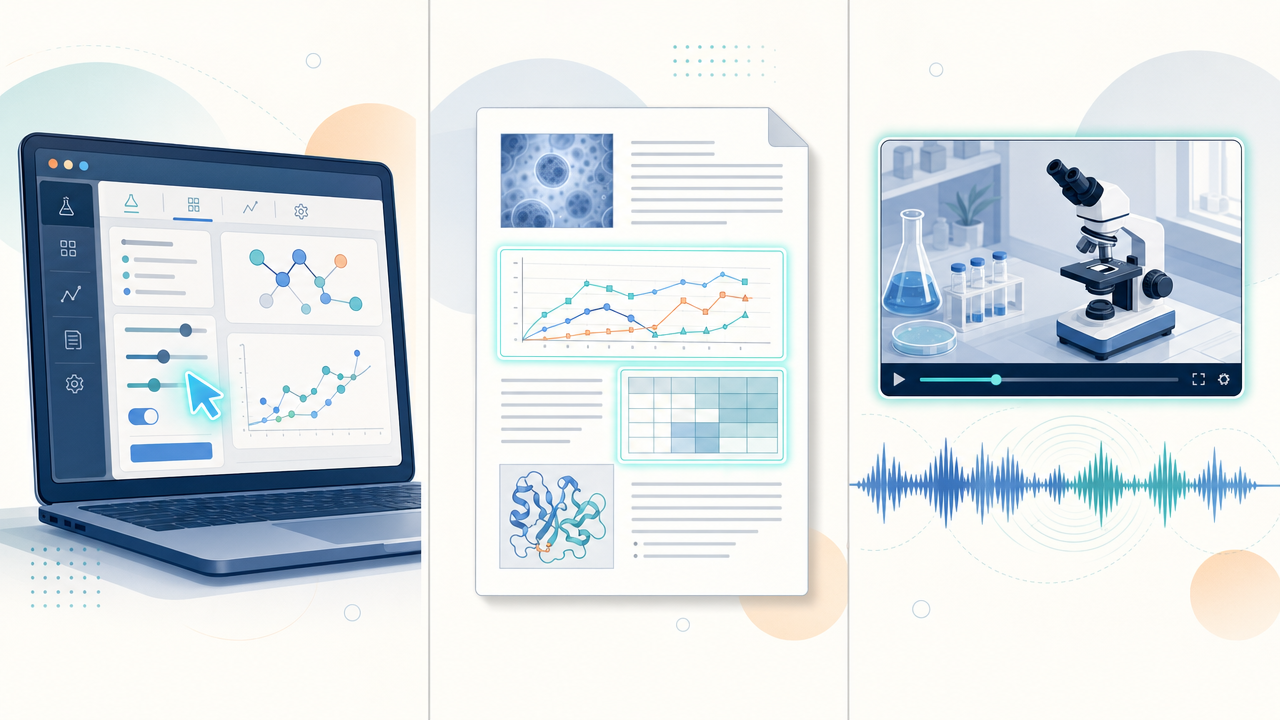

The three application areas highlighted at launch, computer use, document intelligence, and audio-video reasoning, map almost one-to-one onto workflows our customers already run.
NVIDIA highlighted three application areas at launch: computer use, document intelligence, and audio-video reasoning. Each maps directly onto patterns we see thousands of times a week on K-Dense Web.
Computer Use for Instrument and ELN Automation
Scientific software is a graveyard of GUIs. CryoSPARC, Benchling, MNova, GraphPad Prism, the FACS analyzer in the core facility, the LIMS the IRB makes you use, the in-house dashboard one postdoc built in 2017 and nobody has the source for. Most of these tools have no API, or have APIs that lag the GUI by years.
A multimodal agent that can look at a screen, reason about it as a scientist would, and operate the controls turns this from a cost center into an automation surface. Combined with our sandboxed agent runtime built on NVIDIA OpenShell, a Nemotron 3 Nano Omni-powered agent can drive scientific GUIs on a sandboxed Linux desktop and produce reproducible analysis runs without ever touching the underlying credentials.
That is the difference between "AI helped me write the protocol" and "AI ran the analysis end-to-end while I went to seminar."
Document Intelligence on What Scientific Documents Really Are
A scientific paper is not a string of tokens. It is a layout. The figure is the result. The supplementary table is the data. The structure on page 7 is a load-bearing claim. We have been frustrated for a long time that text-first models pretend not to see this.
Nemotron 3 Nano Omni handles charts, tables, screenshots, and mixed-media inputs as first-class evidence. The implications for the workflows we already power on K-Dense Web are direct:
- Competitive intelligence and due diligence. Most of the actual evidence in pharma and biotech sits in slide decks, conference posters, patent figures, and EPA appendices. Multimodal reasoning lets the agent argue against the artifacts, not against summaries of them.
- Regulatory analysis. FDA submissions, EMA responses, ICH guidelines, and clinical trial registries are all heavily figural. Reading the figure correctly is often the entire job.
- Literature synthesis. A figure-aware literature agent can compare your reference micrograph or your target dose-response shape against the figure panels of every candidate paper, not just their captions.
Audio and Video Reasoning Over the Lab Itself
This is the part we are most excited about, and the part we believe is most under-appreciated.
Most lab knowledge never gets written down. It lives in advisor conversations, in lab meeting recordings, in the way someone says "yeah, that's a real band" versus "yeah... that's a band." It lives in the hand gestures during a benchtop demonstration. It lives in the voice memo a clinician dictates between patients.
When the agent can stay in continuous audio-video context across a meeting, a benchtop demonstration, and the resulting micrograph, you start to get something that genuinely behaves like a junior collaborator who was actually in the room. The transcript is no longer a lossy summary the agent has to trust. The audio itself is part of the reasoning.
Open Weights, Local Compute, and Why Both Matter for Science
K-Dense Web is built on a deliberately multi-model architecture. We do not believe one frontier model is going to be the best choice for every step of a research workflow, and we do not want our customers locked to a single provider. Different models have different strengths in coding, in scientific reasoning, in mathematical proof, in visual perception. The platform should route to the right one.
What we look for in a new model is a combination of three things: real capability on workflows our customers actually run, good economics at the volume agentic systems consume, and openness so we and our customers can adapt the model to scientific domains that no general training corpus will ever fully cover. Nemotron 3 Nano Omni hits all three, and the openness piece is the part we want to emphasize most.
Truly Open, Not Just Open Weights
Nemotron 3 Nano Omni is being released, like the rest of the Nemotron 3 family, with open weights, open datasets, and open training techniques. For a scientific platform, that combination is not a nice-to-have. It is structural.
- Open weights mean researchers can host the model themselves, fine-tune it on proprietary data, and pin a specific checkpoint forever. A reviewer in 2030 needs to be able to reproduce a 2026 result. You cannot do that against an API endpoint that silently shifts behind you.
- Open datasets mean the model's training distribution is inspectable. For high-stakes scientific use, knowing what was in the corpus is part of validating the agent's behavior, not a footnote.
- Open training techniques mean we, and our customers, can extend the recipe to domains that no general-purpose corpus will ever fully cover: cryo-EM micrographs, mass spectrometry traces, electrophysiology recordings, the specific dialect a research community uses in its slide decks.
That is the difference between "the model knows in general what a Western blot looks like" and "the model knows what a Western blot from your lab's specific protocol looks like, after you fine-tuned it on your past five years of imaging."
Local Compute, From a Desk to a Data Center
Because Nemotron 3 Nano Omni is open and efficient, you can actually run it where your data lives. That is the part of the story we believe is most under-told in AI-for-science.
The model is designed to deploy consistently across the stack: from a DGX Spark on someone's desk or a DGX Station on a lab bench, to an institutional GPU cluster, to a private-cloud VPC, to fully air-gapped environments. Wherever your data has to live, the model can join it. With NVIDIA NeMo for customization and NVIDIA NIM microservices for deployment, the path from a stock checkpoint to a domain-tuned model running on your own infrastructure is a well-paved road, not a research project of its own.
For science, this changes what is even on the table:
- Clinical and patient data. EHR notes, radiology images, genomic variants, ECG traces, and pathology slides often legally cannot leave a hospital network. A multimodal agent that runs inside the institution's firewall can reason over the actual data without anything ever crossing to a third-party API.
- Industrial R&D and IP. A pharma org's compound library, a battery lab's process telemetry, a semiconductor team's defect micrographs. The teams moving fastest with AI right now are the ones who do not have to choose between "use a strong model" and "do not let our crown-jewel data touch someone else's GPUs."
- Sovereignty and data residency. The EU, UK, and a growing list of regions have hard rules about where research data can be processed. Local deployment is the only honest answer for a lot of these constraints.
- Reproducibility. A frozen, locally hosted checkpoint is part of the methods section. The entire weight of computational science depends on results being re-runnable years later. Hosted models, by their nature, drift.
- Economics at agentic scale. Long-running multimodal agents process orders of magnitude more tokens, image frames, and seconds of audio than a chatbot does. Open weights on your own GPUs change the unit economics of running those agents 24/7.
- Compute-to-data, not data-to-compute. Microscopy datasets, sequencing runs, and instrument logs are routinely terabytes per experiment. Bringing the model to the data is fundamentally cheaper and faster than the reverse, and a single DGX Spark next to the instrument is often the right answer.
Continuing a Pattern We Believe In
This pairing also continues a pattern we are happy to be part of. Earlier this year we wrote about pairing NVIDIA OpenShell with Scientific Agent Skills to give research agents a runtime that scientists could actually trust around patient data and HPC credentials. Our optimize-for-gpu skill routinely turns CPU-bound scientific Python into 58x-faster GPU code by leaning on the NVIDIA RAPIDS ecosystem. Nemotron 3 Nano Omni is a natural next step: an open, locally deployable perception layer that lives natively in the same stack the rest of agentic science is converging on.
What's on the Roadmap
We are building a multimodal-first surface in K-Dense Web powered by Nemotron 3 Nano Omni. Some of what is on the immediate roadmap:
- Figure-aware literature search. When you ask K-Dense Web to find papers with a specific kind of survival curve or a specific dose-response shape, the agent compares your reference image against the figure panels of every candidate paper, not just the captions and abstracts.
- Visual due diligence. For competitive intelligence in pharma and biotech, the agent argues against the actual slide decks, posters, and patent figures. Not against summaries of them.
- Voice-native research sessions. Many of our customers are PIs and clinicians who would rather talk through an analysis than type. With audio understanding integrated end-to-end, the conversation itself becomes the orchestration layer for the agent.
- Hands-on instrument control. Combined with our sandboxed agent runtime, the agent can drive scientific GUIs and produce reproducible analysis runs end-to-end.
- Multi-model orchestration. Nemotron 3 Nano Omni handles perception. Larger Nemotron 3 Super and Ultra checkpoints, along with other frontier models from our existing partners, handle complex planning and long-horizon reasoning. The right shape of work goes to the right model.
- Dedicated multimodal Scientific Agent Skills. Our open-source Scientific Agent Skills library already teaches research agents how to do real science across cheminformatics, single-cell biology, structural biology, astronomy, and more. We are extending it with a new generation of skills built specifically for omni-modal models: reading and validating a Western blot, scoring a microscopy panel for QC, turning a clinical Zoom recording into structured trial notes, or operating CryoSPARC's GUI from raw movies to a published reconstruction. Each skill ships as a tested, citeable artifact that scientists can read, audit, and extend.
- K-Dense BYOK powered by local Nemotron 3 Nano Omni. K-Dense BYOK, our MIT-licensed desktop AI co-scientist, will let researchers point their workflows at a locally hosted Nemotron 3 Nano Omni endpoint. You can bring your own API keys for frontier text models when you want them, and run all multimodal perception against a Nemotron 3 Nano Omni instance on your workstation, a lab DGX Spark, or your institution's GPU cluster. Sensitive data never has to leave your network for the agent to see, hear, and reason over it.
- Local K-Dense deployments on DGX Spark and other NVIDIA GPU platforms. We are working with NVIDIA to bring the full K-Dense agentic stack (sandboxed agent runtime, multimodal perception, Scientific Agent Skills, and orchestration) to local NVIDIA GPU platforms including DGX Spark, DGX Station, and on-prem GPU servers. The end state is a research environment where the agents, the models, the skills, and the data all live inside the same trusted boundary, on hardware the institution owns.
There is a longer arc behind all of this. The thesis is simple. AI co-scientists need to see what scientists see. They need to hear what scientists hear. They need to read what scientists read in the form scientists actually read it: figures, tables, slides, screen recordings, voice memos, whiteboards. An agent that has only ever seen text is, on a basic perceptual level, not in the room.
NVIDIA Nemotron 3 Nano Omni is one of the first models we have used that meaningfully closes this gap while remaining open, efficient, and deployable on the researcher's own infrastructure, from a workstation under a bench to an air-gapped institutional cluster. We are proud to be collaborating with NVIDIA on it, and prouder still that the larger story this lands inside, agentic science with eyes, ears, and hands, is happening in the open.
We will share concrete benchmarks, internal evaluations, and our first end-to-end multimodal workflows in K-Dense Web in the coming weeks. If you want to be among the first to try them, reach out.
Ready to bring multimodal agentic science to your lab? Get started on K-Dense Web →
Questions or want to be part of our early multimodal pilot? Join our Slack community or email contact@k-dense.ai.
Related resources:
- NVIDIA's Nemotron 3 Nano Omni announcement
- NVIDIA Nemotron 3 family overview
- The Sandboxed AI Scientist: Pairing NVIDIA OpenShell with Scientific Agent Skills
- GPU-Accelerate Your Science: 58x Average Speedup with a Single Skill
- Agent Skills: The Final Piece for AI-Powered Scientific Research
- K-Dense Web Platform
